On second thought, if you already paid for it, we’ll find some way to use it. PM me when you receive the data and I’ll arrange for some way for you to upload anonymously, whether via email, or dropbox.
Didn’t see this thread until now either. If I order the test can I still submit my sample?
I hav the data already. I will PM you tomorrow when i´m back home. Thank you so much
Sure. Certainly some way for you to submit can be worked out.
No guarantee that further submissions will be used though.
On the verge of analysis of chromosomes 1-22 of v5 data at the moment.
I have my data ready, if its possible to send it to some dropbox or PM?
Sent you a PM with upload instructions.
Thank you.
The most simple form of association analysis of the 23andMe v5-chip data submitted has been completed. This only included autosomes (chromosomes 1-22). As was to be expected, nothing considered statistically significant was found. There was also not much in the way of clustering, which may indicate a genetic locus associated with a disease in question:
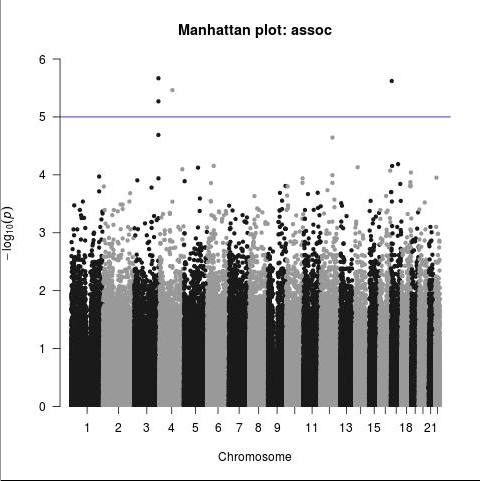
The only SNPs beyond the 1E-5 threshold (still not considered significant at the per-SNP level in GWAS) mapped to 2 genes, P3H2/LEPREL1 and WSCD1, which don’t appear likely to be involved in any mechanism of a post-drug syndrome; either hypothesized mechanisms, or wildly speculated mechanisms. It’s probably a “fluke.” Further analysis will strengthen or weaken this idea.
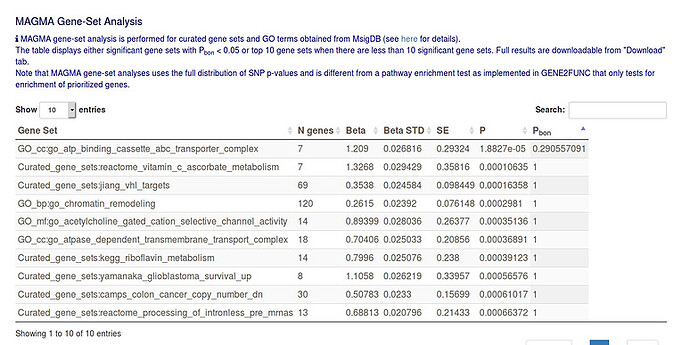
The file containing the results was submitted to a gene ontology service, which attempts to characterize GWAS data according to pathways, functions, or traits, that involve significantly-associated genes. In the absence of significant associations, the service still reports the top-10 pathways, functions, or traits, represented by the data. Basically, the lowest quality setting allowed had to be used for the data to be processed:
Keep in mind that these reported associations were extremely weak and that basing any treatments on something so flimsy is a waste of time and money and could be another risk to your health. Also keep in mind that this is an amateur GWAS study at this point and intended to find something worth approaching professionals with.
There was a bit of a change in plans and the next step will be imputation, which confidently infers the existence of unobserved SNPs based off of observed SNPs in an individual’s genome. Basically, this will provide much more data to work with. Imputation is a computationally intensive process in GWAS, so it may take some time. I’m also committed to some Accutane things in the coming weeks and may not have much time to devote to propeciahelp or this project.
Thanks for the update.
What was the motivation behind the change in plans?
Appreciate the update Dubya, posts are always very informative. It really spells out how complicated this is.
For one, our geneticist friend has expressed his assessment that imputation is needed to sufficiently increase the power of a study with so few genomes. He has given no indication that he is anything but a professional in his field and, as an amateur, I am compelled to give his judgement some weight. I think it’s better to just do the imputation, then analysis, rather than waste time with analysis, imputation, re-analysis.
Also, a forum member who is acquainted with the founder of an imputation service put us in contact with him and he offered to help.
23andMe please.
Nothing against Ancestry, it’s just that our existing data is 23andMe and the 2 companies have used different tests with different chips that included different snps. We chose 23andMe primarily because many members were already genotyped through their service.
Thanks Dubya for the hard work!
1.) Can you explain a little bit more about the top-10 pathways? They are non-significant but show a correlation?
If I see it right “chromatin remodeling” shows the highest correlation?
Though not significant that could be something, not?
There is a connection to epigenetic regulation, senescence and the CHARGE-syndrome that include “micropenis” and “delayed of absent pubertal development”.
If not too much work, it would be nice to see the SNP-name of the top SNPs (to look at them in SNPedia).
2.) Does the data set include the SNPs of the whole 5 isoenzymes of the 5ar-family as well as the SNP of the androgene receptor?:
5α-R1,
5α-R2,
5α-R3,
GPSN2,
GPSN2L
Thanks!
Just ran across this post. Have my 23 and me raw data ready for up load. Am I able to submit?
@Akiyah, Yes. I will PM you.
@anon5006275, I’ll check again to see if the list of snps associated with chromatin remodeling are accessible. I tried briefly once already. Remember that the association is extremely weak, even for a peripheral gene set.
Also, I’m pretty certain the 5-arI, 5-arII, AR genes, and their flanking regions have already been sequenced in PFS patients in the studies and nothing unusual was found.
In what way are you interested in GPSN2? It is one of the genes whose transcription was repressed ~1.4 fold in a study of isotretinoin (Accutane) treated patients.
@Dubya_B can you check ApoE4 frequency among us ? It gives worst out comes with any kind of brain injury. It is also main risk factor for Late onset Alzheimer’s.
Also HTR2A snp’s might worth checking it is closely related pfc dopamine modulation, memory, pain perception and arousal.
rs6311 – 1438 G/A
The ‘T’ allele has been associated with more receptors/increased gene expression and more active receptors. The ‘ TT’ genotype has been associated with depression, panic disorder, and higher stress response. [71, 72].
The ‘ T’ allele was associated with significant reductions in general health, vitality, and social function [73]. The ‘ T’ allele has also been associated with irritable bowel syndrome (IBS) [44] and chronic fatigue syndrome (CFS) [73].
The ‘ C’ allele is associated with an extraverted personality, Rheumatoid Arthritis and novelty seeking. ‘ CC’ = 3.6x increased risk of sexual dysfunction when taking SSRI Antidepressants. This SNP correlates with rs6313, as in the T allele here will result in the A allele by rs6313 [73].
Oh, wow! GPNS2 is an enzyme with the same function as 5ar. It belongs to the 5ar-family. As they have same structure and functionality it is very likely that also Finasteride will bind to it. The link to Accutane is highly interesting!
Ideally the data set includes also the genes for 5ar-3 - Prof. Zitzmann thinks that is the reason for PFS as he says it is more located in the brain.
GPSN2L should be looked at as well as it also has the same function and structure as 5ar. It belongs to the same isoenzyme family.
Any 23andMe Patient Data Analysis participant who wishes to access the summary results of their imputed genome, please send me a PM and you will be provided with a unique ID and instructions as soon as your data is complete.
The more recent (version-5) chip data is complete as of Feb 27th, with the exception of 1 or 2 genomes that seem to be hung up. The remainder should be available in the next few days.
Hello,
I just received my results from 23andme and I downloaded the raw data, how do I send it to you ?
“Acceptance of new submissions is closed until further notice and basic analysis on the current data set has begun. See the following post in this topic for more details:” oh shit … ok, too bad
Really cutting it close. Will send you a PM so you can supply it via email.
Thanks!
23andMe data can still be supplied by members who have an existing file; however, no one is actively encouraged to go through the trouble of having the test performed for the sake of this GWAS at this point.
Once the imputation has finished, and the second round of analysis has actually begun, it will be too late. There’s no set date of when this will be.