The most simple form of association analysis of the 23andMe v5-chip data submitted has been completed. This only included autosomes (chromosomes 1-22). As was to be expected, nothing considered statistically significant was found. There was also not much in the way of clustering, which may indicate a genetic locus associated with a disease in question:
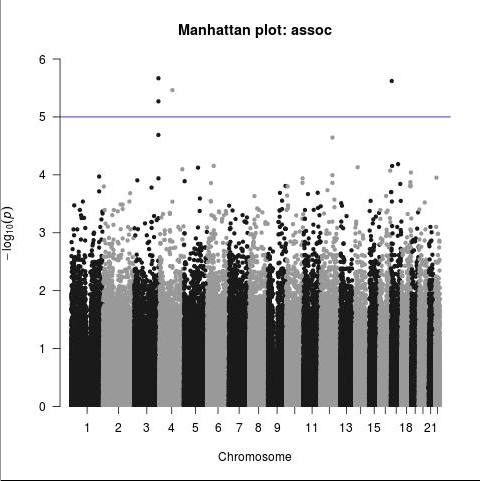
The only SNPs beyond the 1E-5 threshold (still not considered significant at the per-SNP level in GWAS) mapped to 2 genes, P3H2/LEPREL1 and WSCD1, which don’t appear likely to be involved in any mechanism of a post-drug syndrome; either hypothesized mechanisms, or wildly speculated mechanisms. It’s probably a “fluke.” Further analysis will strengthen or weaken this idea.
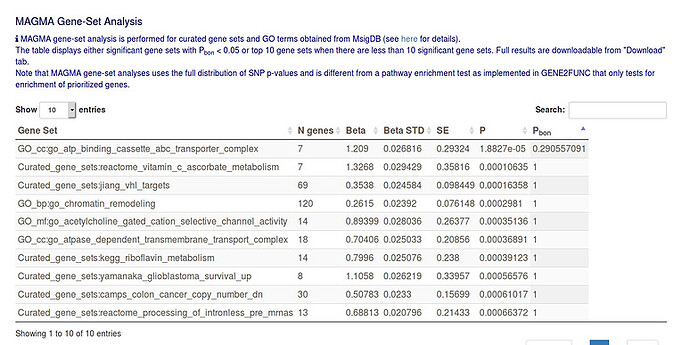
The file containing the results was submitted to a gene ontology service, which attempts to characterize GWAS data according to pathways, functions, or traits, that involve significantly-associated genes. In the absence of significant associations, the service still reports the top-10 pathways, functions, or traits, represented by the data. Basically, the lowest quality setting allowed had to be used for the data to be processed:
Keep in mind that these reported associations were extremely weak and that basing any treatments on something so flimsy is a waste of time and money and could be another risk to your health. Also keep in mind that this is an amateur GWAS study at this point and intended to find something worth approaching professionals with.
There was a bit of a change in plans and the next step will be imputation, which confidently infers the existence of unobserved SNPs based off of observed SNPs in an individual’s genome. Basically, this will provide much more data to work with. Imputation is a computationally intensive process in GWAS, so it may take some time. I’m also committed to some Accutane things in the coming weeks and may not have much time to devote to propeciahelp or this project.